// Research / Computer Vision
Medical AI& imaging computer vision, built as medical device software.
Radiology AI, pathology, ophthalmology, dermatology - DICOM-native pipelines, GPU-resident training that doesn't starve on volumetric data, validation methodology a medical journal would publish, and a deploy posture that runs inside the hospital VLAN. We don't ship a hospital a research notebook.
// What we see
The model is the easy part. The pipeline, the validation, and the audit trail are the work.
01
GPUs starving on a CPU dataloader
Volumetric CT/MRI training pipelines that re-read NIfTI files every epoch, decompress on the CPU, and run 3D augmentation on the CPU end up with GPUs at <30% utilization and an iteration loop measured in days. The model isn't the bottleneck - the data path is.
02
Validation that doesn't survive a second hospital
Most published medical CV results don't generalize across sites. Different scanners, different demographics, different prevalence - and the eval was run on a single-center test set. The first external-validation run is where the AUROC drops 15 points and the customer's regulatory plan unravels.
03
A regulatory pathway retrofitted at audit time
Models trained without frozen weights, without a build provenance record, without per-cohort performance breakdowns, without IEC 62304 traceability. The work to make it audit-ready after the fact is ~3× the work to do it right the first time, and the timeline slips behind the submission window.
// Case Study
Trained a Gleason grading + tumor-segmentation model on full-resolution prostate WSI
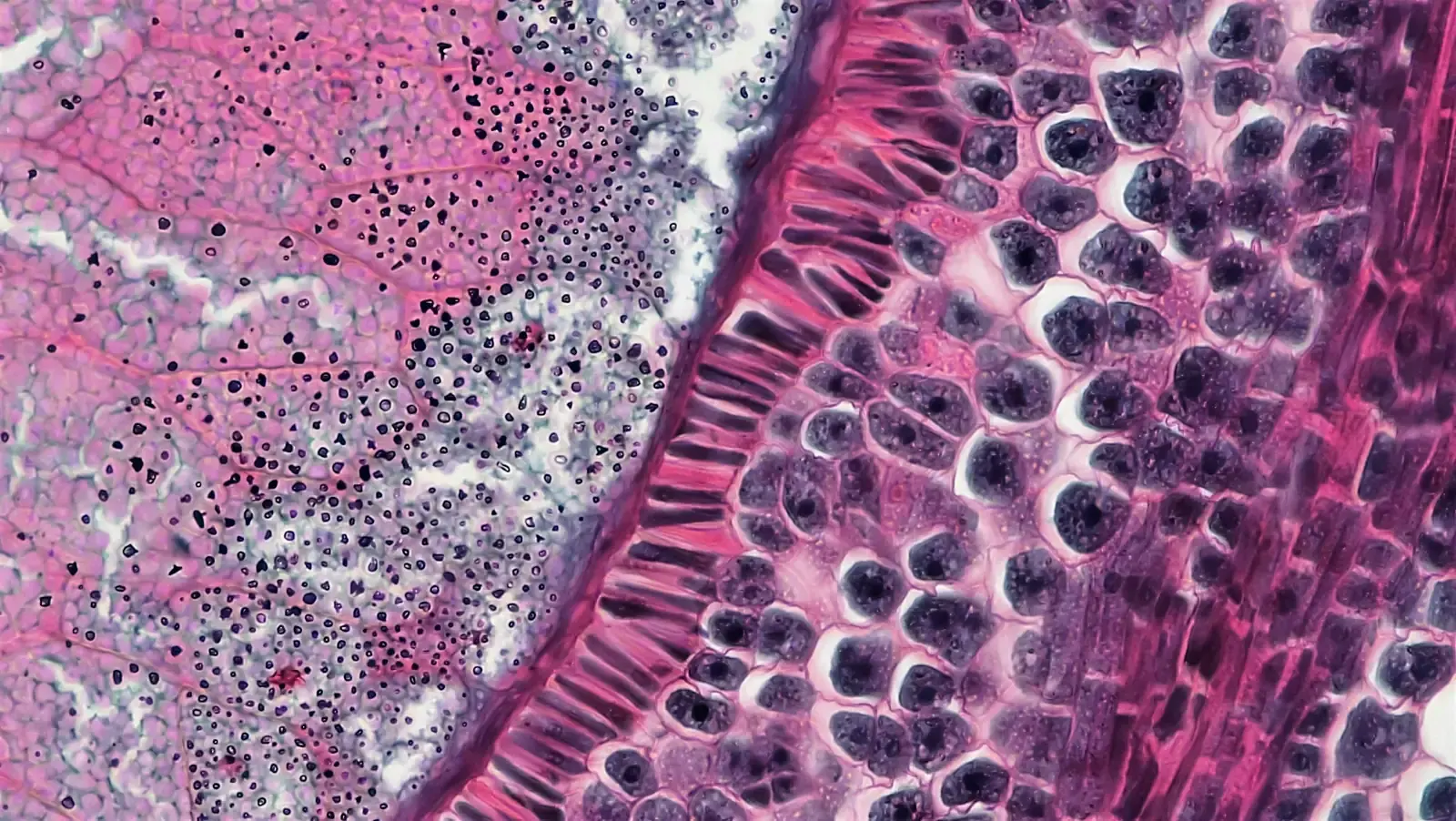
We trained the Gleason grading + tumor-segmentation model - gland-level Gleason pattern classification plus pixel-level cancer segmentation on gigapixel H&E prostate biopsies - for a seed-stage medical imaging startup. The GPUs on their 8×A100 box were idling at <30% under an OpenSlide dataloader and CPU tile augmentation, so we rebuilt the data path GPU-native (Zarr-backed pyramidal tiles, cuCIM preprocessing and stain normalization on-device, Kornia augmentation in the same memory). Single-epoch wallclock collapsed from ~9h to ~50m, GPU utilization went from sub-30% to >90%, and the iteration loop finally caught up with the pathologists' labeling cadence.
9h → ~50m
single-epoch wallclock
>90%
GPU utilization (was <30%)
On-prem
no PHI egress, hospital VLAN
// What we do
Three things that do most of the work.
GPU-native pipelines so the training loop isn't I/O-bound. Validation methodology mapped to the clinical question, not a Kaggle leaderboard. A deploy and audit posture that lands inside the hospital VLAN.
GPU-native data pipelines for volumetric imaging
Zarr-chunked storage instead of NIfTI re-reads, cuCIM for GPU preprocessing, Kornia for 3D augmentation in the same memory. The whole data path lives on the GPU - augmentation is part of the compute graph, not a CPU worker pool the GPUs wait on. Training throughput typically lifts 5–10× on the same hardware.
- Zarr + Blosc/Zstd content-addressed chunked stores, sized to the patch geometry
- cuCIM for HU windowing, isotropic resampling, body-mask, normalization on-device
- Kornia 3D affine + elastic + intensity augmentation in GPU memory, seeded per batch
- Reproducible from run config + dataset hash; validation runs through the same path
Validation methodology that matches the clinical question
We design the validation study before we train. Sensitivity / specificity at clinically meaningful thresholds, prevalence-adjusted PPV, multi-site held-out evaluation, per-cohort breakdowns across demographics and scanners, reader-study comparison against board-certified clinicians. Numbers a medical journal would publish, not a notebook plot.
- Reader studies vs. board-certified specialists with statistical power calculated upfront
- Multi-site held-out evaluation; per-scanner / per-cohort drift reported by default
- Prevalence-adjusted PPV and decision-curve analysis at clinical operating points
- Calibration analysis - Brier score, reliability diagrams, temperature scaling where needed
Hospital-VLAN deploy + SaMD-aligned audit trail
DICOM C-STORE / C-FIND listeners, PACS-integrated reporting via DICOM SR, on-prem GPU appliance, signed install bundles, no PHI egress. Frozen weights with a build provenance record, IEC 62304 lifecycle traceability, and a release process the regulatory team can submit against - built in from day one, not retrofitted.
- DICOM-native ingest and DICOM SR output for PACS-integrated reporting
- On-prem / air-gapped capable: signed bundles, offline registries, no outbound traffic
- Frozen-model release with bit-exact reproducibility and signed weight provenance
- IEC 62304 / ISO 13485 lifecycle traceability the regulatory team can submit with
// Method fit
Medical CV is the right move when the regulatory and clinical bar are part of the brief.
skip it if
You only need a research prototype
If the deliverable is a research result on a public dataset and the path-to-clinical doesn't matter, a single ML engineer with MONAI and a notebook is faster and cheaper than us. Our overhead - validation methodology, audit posture, on-prem deploy - is wasted budget for a notebook.
The use case isn't medical-specific
Generic object detection, OCR on scanned forms, or visual QA on consumer images don't need DICOM, IEC 62304, or reader studies. Use a general computer-vision shop and skip the regulatory tax.
Object DetectionYou can't get clinical data or a clinical site partner
Medical CV without access to real DICOM data and a clinical site for validation is shadow-boxing. We'll help you structure data-sharing agreements and IRB submissions, but if the data path doesn't exist there's nothing to train on and nothing to validate against.
use it if
You're building a clinical product (SaMD or close to it) and the regulatory pathway - FDA 510(k) / De Novo, EU MDR Class IIa/IIb - is part of the brief, not a future problem.
You have, or can get, real clinical data through a hospital partner or research collaboration, and at least one site for external validation.
Deployment is on-prem or in a customer-controlled VPC, and your customers expect clinical data to stay in their network.
// How we work
Validation plan first. GPU-native pipeline second. Then train.
Most failed medical CV engagements failed before the first epoch - validation methodology unclear, data path I/O-bound, regulatory posture retrofitted at the end. We invert the order.
01
Validation plan + clinical question lock
Week one we sit with the clinical lead and the regulatory team and agree on the validation study before we touch a model. Operating points, sensitivity/specificity targets at those points, prevalence assumptions, statistical power, per-cohort breakdowns, the reader-study design. The plan becomes the spec - every model claim afterwards measures against it.
02
GPU-native pipeline + first training run
Zarr build of the curated cohort, cuCIM preprocessing on-device, Kornia 3D augmentation in GPU memory. First training run is set up to be GPU-bound by construction - we measure dataloader latency before we ship a learning-rate sweep. Typical lift: <30% GPU utilization on the customer's old pipeline becomes >90% on the same hardware.
03
Validation, freeze, hand-off with audit trail
Multi-site held-out validation, per-cohort and per-scanner breakdowns, calibration analysis, reader-study comparison. Once the model passes the locked validation plan, weights are frozen, build provenance recorded, and the release bundled for the regulatory team. Hand off the training pipeline (Zarr build script, cuCIM/Kornia config, W&B project, validation harness, retraining runbook) so the team can re-run it as new sites come online.
// Expert insight
“The dirty secret of medical CV is that 80 percent of published results don't generalize across hospitals - different scanners, different demographics, different prevalence. The work that matters is the validation methodology, the data path, and the audit trail. The model architecture is a footnote.”
Karol Gawron
Head of R&D @ bards.ai
// Why bards.ai
Why us, instead of a research lab and good intentions.
Academic publications meet hospital deployments. We bridge the gap between a research result and a clinically usable tool - and we own every layer from DICOM ingest to the regulatory artifact.
Peer-reviewed medical CV background
Founded out of CLARIN-PL academic research. Statistical rigor, validation methodology, and reader-study design are in our DNA - reviewers hold us to the same bar we hold our customer-facing work.
GPU-native pipelines, not CPU dataloaders
Zarr / cuCIM / Kornia for volumetric data is in our standard kit. We've taken customer training pipelines from <30% GPU utilization to >90% on the same hardware - typical lift is 5–10× training throughput, no new GPUs.
DICOM, PACS, HL7 / FHIR fluency
We speak C-STORE, C-FIND, DICOM SR, and the operational language of hospital IT - not just PyTorch. PACS-integrated reporting that lands in the radiologist's worklist instead of a separate console.
On-prem and air-gapped capable
We've deployed CV inside hospital networks that block outbound traffic. Signed install bundles, offline registries, no PHI egress, no cloud dependency in the inference path.
SaMD / IEC 62304 / EU MDR alignment
Frozen weights, build provenance records, lifecycle traceability, per-cohort validation reports. We don't write the 510(k), but every artifact the regulatory team needs to write it ships with the model.
Federated training when data can't move
FedAvg / FedProx with secure aggregation and optional differential privacy, plus per-site domain adaptation for scanner heterogeneity. We've trained across hospitals where data-sharing agreements weren't workable.
// FAQ
Common questions about medical-grade CV
We don't act as the regulatory submitter - that's typically your medical-device team or a specialized RA consultancy. What we do is build every artifact your RA team needs: validation reports, software lifecycle documentation aligned with IEC 62304, risk analysis inputs, and frozen-model release records. We've supported customers through 510(k), De Novo, and EU MDR Class IIa/IIb pathways.
Reader studies against board-certified specialists, multi-site held-out validation, calibration analysis, and per-cohort performance breakdowns. We design the validation study before we train, with statistical power calculated against clinically meaningful effect sizes - so the result is publishable and submission-ready, not a notebook plot.
MONAI's default dataloader is CPU-bound for volumetric data - for many engagements that's fine, for high-volume training on full-resolution CT/MRI it isn't. Zarr gives you constant-time random patch reads and parallel-safe access; cuCIM moves preprocessing onto the GPU; Kornia does the same for 3D augmentation. We use MONAI's transforms when they fit and replace the dataloader when the GPU is starving - it's not an either/or.
We have ongoing collaborations with Polish university hospitals and clinical research centers, and we work with customers' clinical-site networks for cross-site validation. For greenfield projects without an existing site network, we help structure data-sharing agreements and IRB submissions.
Almost always on-prem or in a customer-controlled VPC for clinical deployments. PHI doesn't leave the hospital network. The model runs on customer GPUs (or a small dedicated appliance), integrated with PACS via DICOM listeners. Cloud is a fit only when the customer has already made that compliance decision themselves.
Once a model is validated and submitted, it's frozen - full bit-exact reproducibility, signed weights, signed Docker images, and a build provenance record. Improvements ship as new validated versions through the same release process, not as silent updates. This is non-negotiable for SaMD compliance.
Yes - federated learning is part of our toolkit. We use FedAvg / FedProx with secure aggregation and optional differential privacy, plus per-site domain adaptation to handle scanner heterogeneity. We've done this for cross-hospital training where data-sharing agreements weren't workable.
CT, MRI, X-ray (chest, mammography), pathology whole-slide imaging, OCT, fundus photography, and dermoscopy. The common thread is DICOM-native or vendor-formatted inputs, careful preprocessing, and validation methodology adapted to the modality's prevalence and decision context.
// Let's ship it
Ship CV that holds up to a regulator and a radiologist.
Tell us your modality, your clinical use case, and your target pathway. We'll come back with a validation plan, a pipeline design, and a delivery roadmap - usually within a business day.
Karol Gawron
Head of R&D @ bards.ai