01 · GPUs idle at sub-30% while the data loader does all the work
The Challenge
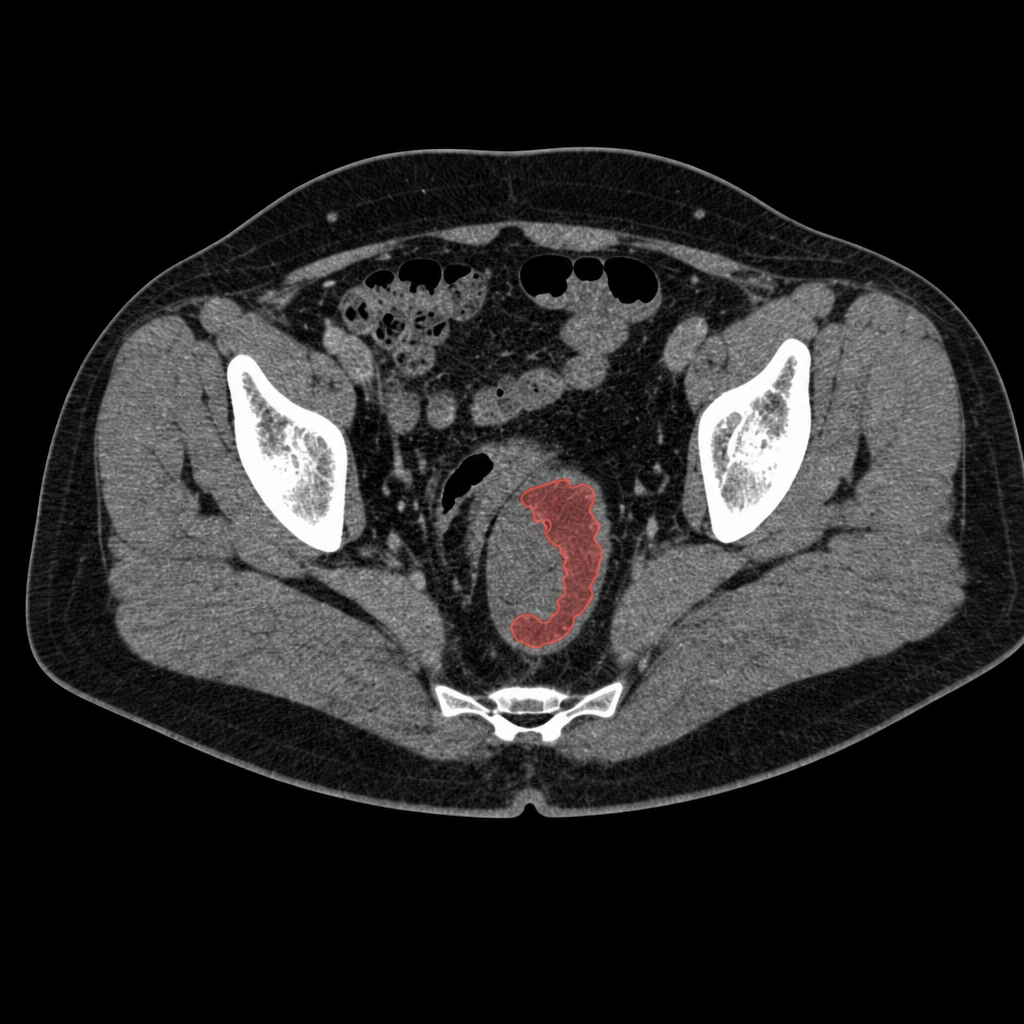
The customer is an early-stage medical imaging startup. The model they needed to ship was a single-head detection + segmentation network on full-resolution abdominal CT - lesion-level localization (colon polyps, suspicious wall thickening, primary-tumor candidates) and voxel-level segmentation of the same lesions, so a downstream radiologist gets both a flag and a contour they can review in the PACS viewer. The clinical promise is real: CT colonography and contrast-enhanced abdominal CT are routine, and a second-reader AI that surfaces missed lesions with a localized mask has a credible path to clinical adoption. The problem was that the team's training loop wasn't keeping up with their own labeling throughput.
A single annotated CT volume in their cohort is ~512×512×800 voxels at sub-millimeter spacing - typically 0.5–2 GB on disk per study, with multiple series (arterial, portal-venous, delayed) per patient. Their training pipeline read NIfTI files end-to-end on every epoch, decompressed on the CPU, ran 3D augmentations (random crops, elastic deformations, intensity jitter, rotations) on the CPU, and then pushed the result to the GPU. The result on their 8×A100 box was textbook starvation: GPU utilization hovering around 25%, single-epoch wallclock around 9 hours on the curated training set, and an iteration loop where a single hyperparameter sweep took most of a week.
Two non-negotiables shaped the rebuild. PHI couldn't leave the hospital VLAN - DICOM ingest, training data, and model artifacts all had to live inside the customer-controlled environment, with no cloud step in the loop. And the rebuild had to land on the same 8×A100 box the customer already owned; raising more capital to buy more GPUs because the data loader was inefficient wasn't a plan their seed round could afford.